Creating a Lizard Brain Atlas
Until recently, reptilian evolutionary studies lacked an important resource – a lizard brain atlas. As the subject of numerous ecological and behavioural studies, the Australian tawny dragon (Agamidae: Ctenophorus decresii) was an ideal candidate for creating a high-resolution MRI atlas of a representative scaled reptile (squamata). Such data is not only a resource for studies of the genus but also informs environmental decision making through an improved understanding of animal adaptation and evolution.
In an impressive cross-disciplinary collaboration, then-PhD candidate Dr Daniel Hoops worked with a team of scientists from Spain, Canada, NSW, VIC, ACT, and QLD to explore brain evolution in reptiles.
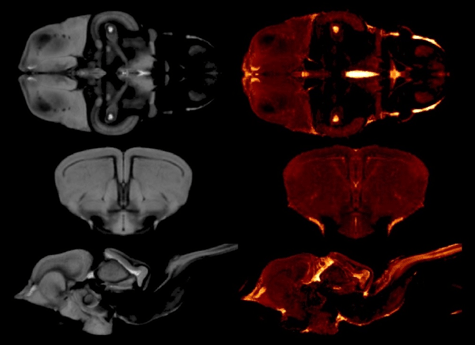
All brains were scanned at the NCRIS-enabled Western Sydney University (WSU) NIF node on the Bruker 11.7 T MRI scanner. Hundreds of brains of a number of species of Australian tawny dragons of both genders were scanned at the WSU NIF Node at 100 μm resolution. To largely automate the measurement of the volumes of the six subdivisions in each of the models, model-based segmentation was used. First, a global model of the tawny dragon brain was created, and the six subdivisions were traced, drawing on expertise at the NCRIS-enabled QLD NIF node. Tracing was achieved first by using the interpolation function of the program Amira (Fischer 3D), followed by manual correction of every slice in all 3 dimensions, resulting in one 3D label for each subdivision of the tawny dragon MRI model (this process is called manual segmentation).
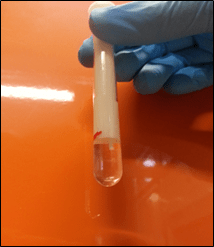
The 3D printed lizard brain holder designed and printed at the WSU node. The brains are buoyant in Fomblin (a dense fluorinated lubricant with no protons and hence no NMR/MRI signal). The Fomblin eliminates the image artefacts that would otherwise occur due to susceptibility problems from the brain and air interface. The 3D printed brain holder secures the brain in place (the brain stem is just visible in the image). Due to the precise placement enabled by the brain holder, tuning and matching and shimming were only needed for the first brain in a run permitting a throughput of four brains an hour each scanned in 3D at 100 μm isotropic resolution.
There being no neuroanatomical atlas available for an agamid brain required the use of numerous available atlases for the brains of other lizards to delineate the six subdivisions accurately. This presented an opportunity to create our own atlas, and thirteen tawny dragon brains were scanned at a higher 50 μm isotropic resolution. Using the informatics expertise at the QLD NIF node, these images were processed using minimum-deformation modelling to create a 20 μm isotropic atlas.
There is a wealth of ecological data to analyse with brain data. The atlas allows a better comprehension of brain areas (nuclei, fibre tracts) and is a valuable resource for future comparative evolution imaging studies, both on the same and other species. The work also provided a novel application—testing and potentially informing further development—of the minimum-deformation modelling technique used to create the 20 μm isotropic super-resolution dataset from the thirteen 50 μm reptilian brain datasets.
Evidence of concerted brain evolution (brain regions co-evolving) was found for body size and mosaic brain evolution (brain regions evolving independently) for ecological habitat. Furthermore, the MRI atlas was a success and has become a useful resource for Agamid evolutionary studies and evolutionary studies in general. The MRI model and atlas of the tawny dragon brain can be viewed online and downloaded using the Wiley Biolucida Server at wiley.biolucida.net.
Further reading:
Publications
- A 3D MRI-based atlas of a lizard brain – 10.1002/cne.24480
- Evidence for concerted and mosaic brain evolution in dragon lizards – 10.1159/000478738
- Sexual selection predicts brain structure in dragon lizards – 10.1111/jeb.12984
- The Secret Caverns of the Dragon’s Brain: Current and Potential Contributions of Lizards to Evolutionary Neuroscience – 10.1159/000486529
Daniel Hoop’s PhD Thesis
- Structure and Evolution of Dragon Brains – 10.25911/5d778b01ca51d
Atlas and data sets
This story was contributed by Dr Tim Stait-Gardner and Dr Daniel Hoops, formerly at ANU, Australia, now at McGill University, Canada.



